CPS-ZJU scientists develop molecular docking technology based on deep learning
2023-09-23 | 药学院英文网
In September 2023, the team of Professor Tingjun Hou, Professor Changyu Xie, Researcher Peichen Pan of from the College of Pharmaceutical Sciences of Zhejiang University , and Researcher Guangyong Chen from Zhejiang Lab jointly published a paper titled "Efficient and accurate large library ligand docking with KarmaDock" in Nature Computational Science. In this paper, a deep learning model called KarmaDock is proposed to generate binding conformations and predict binding strengths quickly and accurately. This model is aligned by post-processing techniques such as force field optimisation and RDkit conformation alignment to correct the bond length and angle inaccuracies detected in the predicted ligand conformation. The method is at least 163 times faster and at least 22.3% more accurate than traditional methods. KarmaDock completed the virtual screening of the ChemDiv+Specs database (1.77 million molecules in total) against the LTK target in just 8.4h, and found over a dozen experimentally-validated active compounds amongst the screened compounds.

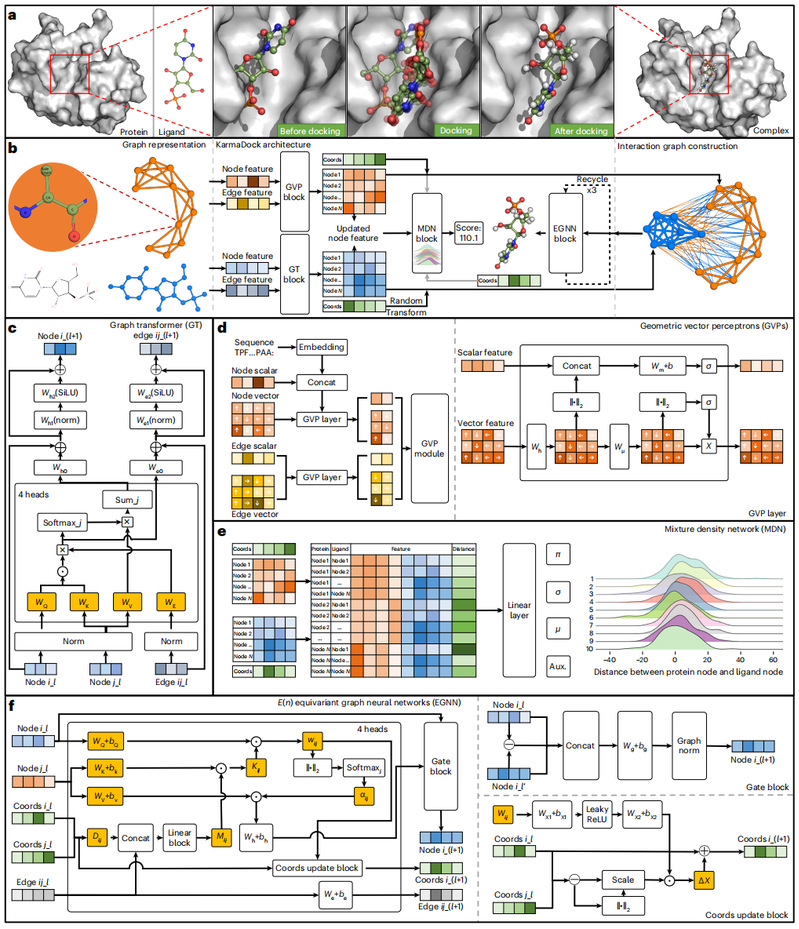
Ligand docking is one of the core technologies in structure-based virtual screening for drug discovery. However, conventional docking tools and existing deep learning tools may suffer from limited performance in terms of speed, pose quality and binding affinity accuracy. Here we propose KarmaDock, a deep learning approach for ligand docking that integrates the functions of docking acceleration, binding pose generation and correction, and binding strength estimation. The three-stage model consists of the following components: (1) encoders for the protein and ligand to learn the representations of intramolecular interactions; (2) E(n) equivariant graph neural networks with self-attention to update the ligand pose based on both protein–ligand and intramolecular interactions, followed by postprocessing to ensure chemically plausible structures; (3) a mixture density network for scoring the binding strength. KarmaDock was validated on four benchmark datasets and tested in a real-world virtual screening project that successfully identified experiment-validated active inhibitors of leukocyte tyrosine kinase (LTK).
Xujun Zhang and Haotian Zhang (Ph.D Candidates) are the co-first authors. Professor Tingjun Hou, Professor Changyu Xie, Researcher Peichen Pan of Zhejiang University and Researcher Guangyong Chen from Zhejiang Lab are the co-corresponding authors.
Link: https://www.nature.com/articles/s43588-023-00511-5
Translator: Zesheng Cheng
Editor: Yichen Zhu
NEWS
-
20
2026.04
-
30
2026.03
-
10
2025.12
-
27
2025.11
-
25
2025.11
-
03
2025.11
